Flow cytometry in oral cytology: Improved brush biopsy-based delineation of oral malignant and potentially malignant lesions

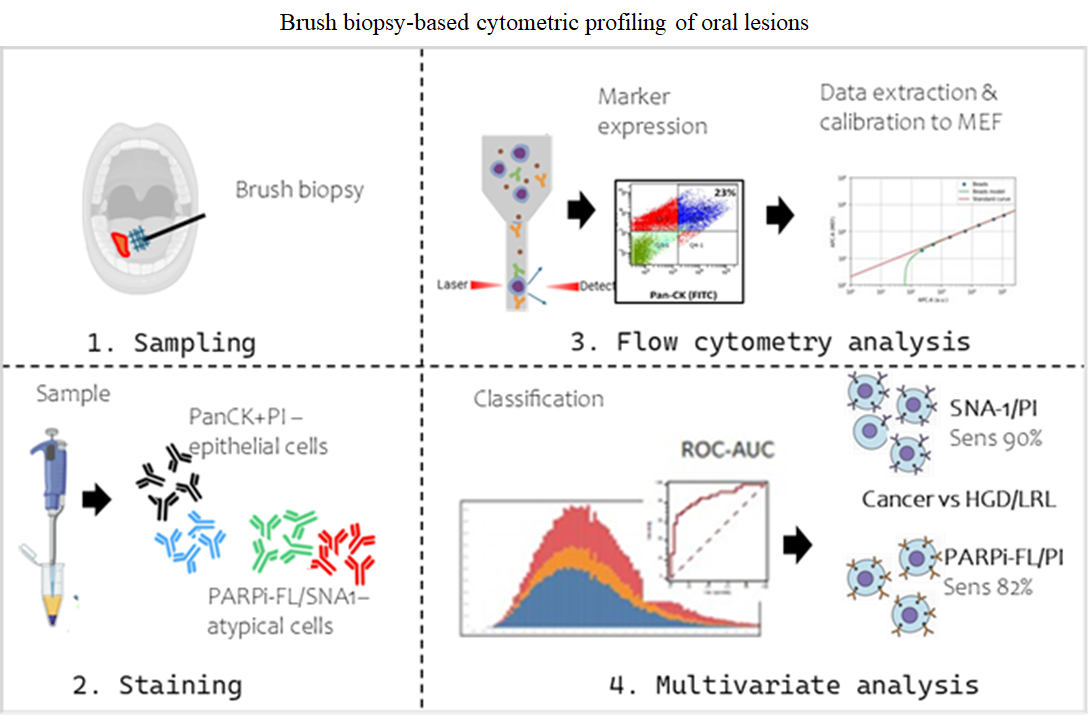
Background: Brush biopsy is a minimally invasive method for early detection of oral squamous cell carcinoma (OSCC). Enhanced accuracy for clinical utility depends on analysis of the whole cell population and automated cohort classifications. Aim: This study aims to delineate OSCC, high-grade dysplasia (HGD), and low-risk lesions (LRLs) by profiling single-cell level alterations using multiplexed flow cytometry. Methods: Brush-biopsy samples were analyzed from patients with LRL, HGD, and OSCC. Flow cytometry analysis was standardized to ascertain cell distribution, heterogeneity, and epithelial cell content. Markers were used for epithelial cell (Pan-Cytokeratin [Pan-CK]/ propidium iodide [PI]) and atypical cell (Sambucus–Nigra–Agglutinin-1 [SNA-1]/ polyadenosine diphosphate-ribose polymerase inhibitor [PARPi-FL]) delineation. In addition, scatter properties and molecular-equivalence fluorescence (MEF) values of markers were analyzed for cohort classification. Results: Brush-biopsy samples from OSCC/HGD patients showed heterogeneity in the percentage of Pan-CK+ve/PI+ve cells. Significant variation in MEF values of SNA-1/PARPi-FL/PI delineated the OSCC cohort (area under the curve > 0.85). Furthermore, the markers in combination with scatter properties delineated OSCC (multivariate logistic regression; sensitivity: 90%, specificity: 82%). The analysis of the forward-scatter height-to-area ratio delineated HGD from low-risk lesions by capturing the morphology-based cellular differences. Conclusions: These results suggest that a flow cytometry-based analysis of brushbiopsy samples may serve as an adjunct tool for risk stratification of oral lesions. Relevance for patients: This study provides evidence towards the application of flow cytometry as an objective, quantitative adjunct to conventional cytology, and improves early detection and risk stratification of oral lesions using a minimally invasive sampling method, thereby supporting timely clinical decision-making and patient management.
- Messadi DV. Diagnostic aids for detection of oral precancerous conditions. Int J Oral Sci. 2013;5(2):59-65. doi: 10.1038/ijos.2013.24
- Giovannacci I, Vescovi P, Manfredi M, Meleti M. Noninvasive visual tools for diagnosis of oral cancer and dysplasia: A systematic review. Med Oral Patol Oral Cir Bucal. 2016;21(3):e305-e315. doi: 10.4317/medoral.20996
- Fedele S. Diagnostic aids in the screening of oral cancer. Head Neck Oncol. 2009;1:5. doi: 10.1186/1758-3284-1-5
- Mendonca P, Sunny SP, Mohan U, Birur NP, Suresh A, Kuriakose MA. Non-invasive imaging of oral potentially malignant and malignant lesions: A systematic review and meta-analysis. Oral Oncol. 2022;130:105877. doi: 10.1016/j.oraloncology.2022.105877
- Demetrio de Souza Franca P, Kossatz S, Brand C, et al. A phase I study of a PARP1-targeted topical fluorophore for the detection of oral cancer. Eur J Nucl Med Mol Imaging. 2021;48(11):3618-3630. doi: 10.1007/s00259-021-05372-6
- Kossatz S, Pirovano G, Demetrio De Souza Franca P, et al. Validation of the use of a fluorescent PARP1 inhibitor for the detection of oral, oropharyngeal and oesophageal epithelial cancers. Nat Biomed Eng. 2020;4(3):272-285. doi: 10.1038/s41551-020-0526-9
- Kumari P, Debta P, Dixit A. Oral Potentially Malignant Disorders: Etiology, Pathogenesis, and Transformation Into Oral Cancer. Front Pharmacol. 2022;13:825266. doi: 10.3389/fphar.2022.825266
- Felthaus O, Ettl T, Gosau M, et al. Cancer stem cell-like cells from a single cell of oral squamous carcinoma cell lines. Biochem Biophys Res Commun. 2011;407(1):28-33. doi: 10.1016/j.bbrc.2011.02.084
- Patel SS, Shah KA, Shah MJ, Kothari KC, Rawal RM. Cancer stem cells and stemness markers in oral squamous cell carcinomas. Asian Pac J Cancer Prev. 2014;15(20):8549- 8556. doi: 10.7314/apjcp.2014.15.20.8549
- Laprise C, Shahul HP, Madathil SA, et al. Periodontal diseases and risk of oral cancer in Southern India: Results from the HeNCe Life study. Int J Cancer. 2016;139(7):1512- 1519. doi: 10.1002/ijc.30201
- Idrees M, Farah CS, Sloan P, Kujan O. Oral brush biopsy using liquid-based cytology is a reliable tool for oral cancer screening: A cost-utility analysis: Oral brush biopsy for oral cancer screening. Cancer Cytopathol. 2022;130(9):740-748. doi: 10.1002/cncy.22599
- Sunny SP, D RR, Hariharan A, et al. CD44-SNA1 integrated cytopathology for delineation of high grade dysplastic and neoplastic oral lesions. PLoS One. 2023;18(9):e0291972. doi: 10.1371/journal.pone.0291972
- McRae MP, Modak SS, Simmons GW, et al. Point-of-care oral cytology tool for the screening and assessment of potentially malignant oral lesions. Cancer Cytopathol. 2020;128(3):207- 220. doi: 10.1002/cncy.22236
- Dabelsteen E. Cell surface carbohydrates as prognostic markers in human carcinomas. J Pathol. 1996;179(4):358- 369. doi: 10.1002/(SICI)1096-9896(199608)179:4<358::AIDPATH564>3.0.CO;2-T
- Vigneswaran N, Peters KP, Hornstein OP, Diepgen TL. Alteration of cell surface carbohydrates associated with ordered and disordered proliferation of oral epithelia: a lectin histochemical study in oral leukoplakias, papillomas and carcinomas. Cell Tissue Kinet. 1990;23(1):41-55. doi: 10.1111/j.1365-2184.1990.tb01108.x
- Pinho SS, Reis CA. Glycosylation in cancer: mechanisms and clinical implications. Nat Rev Cancer. 2015;15(9):540- 555. doi: 10.1038/nrc3982
- Strome A, Kossatz S, Zanoni DK, Rajadhyaksha M, Patel S, Reiner T. Current Practice and Emerging Molecular Imaging Technologies in Oral Cancer Screening. Mol Imaging. 2018;17:1536012118808644. doi: 10.1177/1536012118808644
- Mazumdar S, SenGupta SK, Param R, Sinha SN. Binding pattern of eight different lectins in healthy subjects and patients with dysplastic and malignant lesions of the oral cavity. Int J Oral Maxillofac Surg. 1993;22(5):301-305. doi: 10.1016/s0901-5027(05)80521-1
- Pillai KR, Remani P, Kannan S, et al. Lectin histochemistry of oral premalignant and malignant lesions: correlation of JFL and PNA binding pattern with tumour progression. Eur J Cancer B Oral Oncol. 1996;32B(1):32-37. doi: 10.1016/0964-1955(95)00051-8
- Li Y, Tong Y, Liu J, Lou J. The Role of MicroRNA in DNA Damage Response. Front Genet. 2022;13:850038. doi: 10.3389/fgene.2022.850038
- Wang Y, Taniguchi T. MicroRNAs and DNA damage response: implications for cancer therapy. Cell Cycle. 2013;12(1):32-42. doi: 10.4161/cc.23051
- Lajer CB, Garnaes E, Friis-Hansen L, et al. The role of miRNAs in human papilloma virus (HPV)-associated cancers: bridging between HPV-related head and neck cancer and cervical cancer. Br J Cancer. 2012;106(9):1526- 1534. doi: 10.1038/bjc.2012.109
- Manikandan M, Deva Magendhra Rao AK, Arunkumar G, et al. Oral squamous cell carcinoma: microRNA expression profiling and integrative analyses for elucidation of tumourigenesis mechanism. Mol Cancer. 2016;15(1):28. doi: 10.1186/s12943-016-0512-8
- Agrawal P, Kurcon T, Pilobello KT, et al. Mapping posttranscriptional regulation of the human glycome uncovers microRNA defining the glycocode. Proc Natl Acad Sci USA. 2014;111(11):4338-4343. doi: 10.1073/pnas.1321524111
- Ibrahim SA, Hassan H, Gotte M. MicroRNA regulation of proteoglycan function in cancer. FEBS J. 2014;281(22):5009- 5022. doi: 10.1111/febs.13026
- Thomas D, Rathinavel AK, Radhakrishnan P. Altered glycosylation in cancer: A promising target for biomarkers and therapeutics. Biochim Biophys Acta Rev Cancer. 2021;1875(1):188464. doi: 10.1016/j.bbcan.2020.188464
- Kossatz S, Weber W, Reiner T. Detection and Delineation of Oral Cancer With a PARP1-Targeted Optical Imaging Agent. Mol Imaging. 2017;16:1536012117723786. doi: 10.1177/1536012117723786
- Wang F, Gouttia OG, Wang L, Peng A. PARP1 Upregulation in Recurrent Oral Cancer and Treatment Resistance. Front Cell Dev Biol. 2021;9:804962. doi: 10.3389/fcell.2021.804962
- Sunny SP, Khan AI, Rangarajan M, et al. Oral epithelial cell segmentation from fluorescent multichannel cytology images using deep learning. Comput Methods Programs Biomed. 2022;227:107205. doi: 10.1016/j.cmpb.2022.107205
- Mello FW, Melo G, Guerra ENS, Warnakulasuriya S, Garnis C, Rivero ERC. Oral potentially malignant disorders: A scoping review of prognostic biomarkers. Crit Rev Oncol Hematol. 2020;153:102986. doi: 10.1016/j.critrevonc.2020.102986
- Scott IS, Odell E, Chatrath P, et al. A minimally invasive immunocytochemical approach to early detection of oral squamous cell carcinoma and dysplasia. Br J Cancer. 2006;94(8):1170-1175. doi: 10.1038/sj.bjc.6603066
- Datta M, Laronde D, Palcic B, Guillaud M. The role of DNA image cytometry in screening oral potentially malignant lesions using brushings: A systematic review. Oral Oncol. 2019;96:51-59. doi: 10.1016/j.oraloncology.2019.07.006
- McKinnon KM. Flow Cytometry: An Overview. Curr Protoc Immunol. 2018;120:5.1.1-5.1.11. doi: 10.1002/cpim.40
- Barnett D, Walker B, Landay A, Denny TN. CD4 immunophenotyping in HIV infection. Nat Rev Microbiol. 2008;6(11 Suppl):S7-S15. doi: 10.1038/nrmicro1998
- Grewal RK, Chetty M, Abayomi EA, Tomuleasa C, Fromm JR. Use of flow cytometry in the phenotypic diagnosis of hodgkin’s lymphoma. Cytometry B Clin Cytom. 2019;96(2):116-127. doi: 10.1002/cyto.b.21724
- Gaur G, Awasthi NP, Gupta A, et al. Diagnostic accuracy of flow cytometry in detecting malignant epithelial cells in serous effusions. J Am Soc Cytopathol. 2023;12(6):423-435. doi: 10.1016/j.jasc.2023.09.003
- Chen CC, Chi CY. Biosafety in the preparation and processing of cytology specimens with potential coronavirus (COVID-19) infection: Perspectives from Taiwan. Cancer Cytopathol. 2020;128(5):309-316. doi: 10.1002/cncy.22280
- Straccia P, Rossi ED, Martini M, et al. Description of a new biosafe procedure for cytological specimens from patients with COVID-19 processed by liquid-based preparations. Cancer Cytopathol. 2020;128(12):905-909. doi: 10.1002/cncy.22341
- Birur PN, Patrick S, Warnakulasuriya S, et al. Consensus guidelines on management of oral potentially malignant disorders. Indian J Cancer. 2022;59(3):442-453. doi: 10.4103/ijc.IJC_128_21
- Khan AS, Khan ZA, Nisar M, et al. Description of clinicopathological characteristics of oral potentially malignant disorders with special focus on two histopathologic grading systems and subepithelial inflammatory infiltrate. J Cancer Res Ther. 2023;19(Suppl 2):S724-S730. doi: 10.4103/jcrt.jcrt_969_22
- Sunny S, Baby A, James BL, et al. A smart tele-cytology point-of-care platform for oral cancer screening. PLoS One. 2019;14(11):e0224885. doi: 10.1371/journal.pone.0224885
- Lopresti A, Malergue F, Bertucci F, et al. Sensitive and easy screening for circulating tumor cells by flow cytometry. JCI Insight. 2019;4(14):e128180. doi: 10.1172/jci.insight.128180
- Drescher H, Weiskirchen S, Weiskirchen R. Flow Cytometry: A Blessing and a Curse. Biomedicines. 2021;9(11):1613. doi: 10.3390/biomedicines9111613
- Finak G, Langweiler M, Jaimes M, et al. Standardizing Flow Cytometry Immunophenotyping Analysis from the Human ImmunoPhenotyping Consortium. Sci Rep. 2016;6:20686. doi: 10.1038/srep20686
- Tadijan A, Humphries JD, Samarzija I, et al. The Tongue Squamous Carcinoma Cell Line Cal27 Primarily Employs Integrin alpha6beta4-Containing Type II Hemidesmosomes for Adhesion Which Contribute to Anticancer Drug Sensitivity. Front Cell Dev Biol. 2021;9:786758. doi: 10.3389/fcell.2021.786758
- Kossatz S, Brand C, Gutiontov S, et al. Detection and delineation of oral cancer with a PARP1 targeted optical imaging agent. Sci Rep. 2016;6:21371. doi: 10.1038/srep21371
- Castillo-Hair SM, Sexton JT, Landry BP, Olson EJ, Igoshin OA, Tabor JJ. FlowCal: A User-Friendly, Open Source Software Tool for Automatically Converting Flow Cytometry Data from Arbitrary to Calibrated Units. ACS Synth Biol. 2016;5(7):774-780. doi: 10.1021/acssynbio.5b00284
- Le Lann L, Jouve PE, Alarcon-Riquelme M, et al. Standardization procedure for flow cytometry data harmonization in prospective multicenter studies. Sci Rep. 2020;10(1):11567. doi: 10.1038/s41598-020-68468-3
- Lehmann R. 3σ-Rule for Outlier Detection from the Viewpoint of Geodetic Adjustment. J Surv Eng. 2013;139(4):157-165. doi: 10.1061/(asce)su.1943-5428.0000112
- Kanamori E, Itoh M, Tojo N, Koyama T, Nara N, Tohda S. Flow cytometric analysis of Notch1 and Jagged1 expression in normal blood cells and leukemia cells. Exp Ther Med. 2012;4(3):397-400. doi: 10.3892/etm.2012.633
- Gonzalez-Moles MA, Aguilar-Ruiz M, Ramos-Garcia P. Challenges in the Early Diagnosis of Oral Cancer, Evidence Gaps and Strategies for Improvement: A Scoping Review of Systematic Reviews. Cancers (Basel). 2022;14(19):4967. doi: 10.3390/cancers14194967
- Mittal M, Siddiqui MR, Tran K, Reddy SP, Malik AB. Reactive oxygen species in inflammation and tissue injury. Antioxid Redox Signal. 2014;20(7):1126-1167. doi: 10.1089/ars.2012.5149
- Rembialkowska N, Kocik Z, Klosinska A, et al. Inflammation- Driven Genomic Instability: A Pathway to Cancer Development and Therapy Resistance. Pharmaceuticals (Basel). 2025;18(9):1406. doi: 10.3390/ph18091406
- Huang R, Zhou PK. DNA damage repair: historical perspectives, mechanistic pathways and clinical translation for targeted cancer therapy. Signal Transduct Target Ther. 2021;6(1):254. doi: 10.1038/s41392-021-00648-7
- Pillitteri LJ, Guo X, Dong J. Asymmetric cell division in plants: mechanisms of symmetry breaking and cell fate determination. Cell Mol Life Sci. 2016;73(22):4213-4229. doi: 10.1007/s00018-016-2290-2
- Mohanta A, Mohanty PK, Parida G. Cytomorphometric analysis of keratinized round cells in human oral carcinoma. J Cytol. 2015;32(2):107-112. doi: 10.4103/0970-9371.160561
- Remmerbach TW, Meyer-Ebrecht D, Aach T, et al. Toward a multimodal cell analysis of brush biopsies for the early detection of oral cancer. Cancer. 2009;117(3):228-235. doi: 10.1002/cncy.20028
- Peters JM, Ansari MQ. Multiparameter flow cytometry in the diagnosis and management of acute leukemia. Arch Pathol Lab Med. 2011;135(1):44-54. doi: 10.5858/2010-0387-RAR.1
- Betters DM. Use of Flow Cytometry in Clinical Practice. J Adv Pract Oncol. 2015;6(5):435-440. doi: 10.6004/jadpro.2015.6.5.4
- Bockerstett KA, Wong CF, Koehm S, Ford EL, DiPaolo RJ. Molecular Characterization of Gastric Epithelial Cells Using Flow Cytometry. Int J Mol Sci. 2018;19(4):1096. doi: 10.3390/ijms19041096
- Tinnevelt GH, Kokla M, Hilvering B, et al. Novel data analysis method for multicolour flow cytometry links variability of multiple markers on single cells to a clinical phenotype. Sci Rep. 2017;7(1):5471. doi: 10.1038/s41598-017-05714-1
- Schmid I, Dagarag MD, Hausner MA, et al. Simultaneous flow cytometric analysis of two cell surface markers, telomere length, and DNA content. Cytometry. 2002;49(3):96-105. doi: 10.1002/cyto.10163
- Gough A, Stern AM, Maier J, et al. Biologically Relevant Heterogeneity: Metrics and Practical Insights. SLAS Discov. 2017;22(3):213-237. doi: 10.1177/2472555216682725
- Diaz-Cano SJ. Tumor heterogeneity: mechanisms and bases for a reliable application of molecular marker design. Int J Mol Sci. 2012;13(2):1951-2011. doi: 10.3390/ijms13021951
- Parshenkov A, Hennet T. Glycosylation-Dependent Induction of Programmed Cell Death in Murine Adenocarcinoma Cells. Front Immunol. 2022;13:797759. doi: 10.3389/fimmu.2022.797759
- Kurkalang S, Roy S, Acharya A, et al. Single-cell transcriptomic analysis of gingivo-buccal oral cancer reveals two dominant cellular programs. Cancer Sci. 2023;114(12):4732-4746. doi: 10.1111/cas.15979
- Fedorec AJH, Robinson CM, Wen KY, Barnes CP. FlopR: An Open Source Software Package for Calibration and Normalization of Plate Reader and Flow Cytometry Data. ACS Synth Biol. 2020;9(9):2258-2266. doi: 10.1021/acssynbio.0c00296
- Rutter JW, Ozdemir T, Galimov ER, et al. Detecting Changes in the Caenorhabditis elegans Intestinal Environment Using an Engineered Bacterial Biosensor. ACS Synth Biol. 2019;8(12):2620-2628. doi: 10.1021/acssynbio.9b00166
- Beal J, Overney C, Adler A, Yaman F, Tiberio L, Samineni M. TASBE Flow Analytics: A Package for Calibrated Flow Cytometry Analysis. ACS Synth Biol. 2019;8(7):1524-1529. doi: 10.1021/acssynbio.8b00533
- Jara-Lazaro AR, Thamboo TP, Teh M, Tan PH. Digital pathology: exploring its applications in diagnostic surgical pathology practice. Pathology. 2010;42(6):512-518. doi: 10.3109/00313025.2010.508787
- Baeten J, Johnson A, Sunny S, et al. Chairside molecular imaging of aberrant glycosylation in subjects with suspicious oral lesions using fluorescently labeled wheat germ agglutinin. Head Neck. 2018;40(2):292-301. doi: 10.1002/hed.24943
- Sasikumar J, Laha S, Naik B, Das SP. Enhanced visualization of nuclear staining and cell cycle analysis for the human commensal Malassezia. Sci Rep. 2024;14(1):20936. doi: 10.1038/s41598-024-69024-z

