Temporal correlation rough set-based three-way clustering model for chronic kidney disease diagnosis

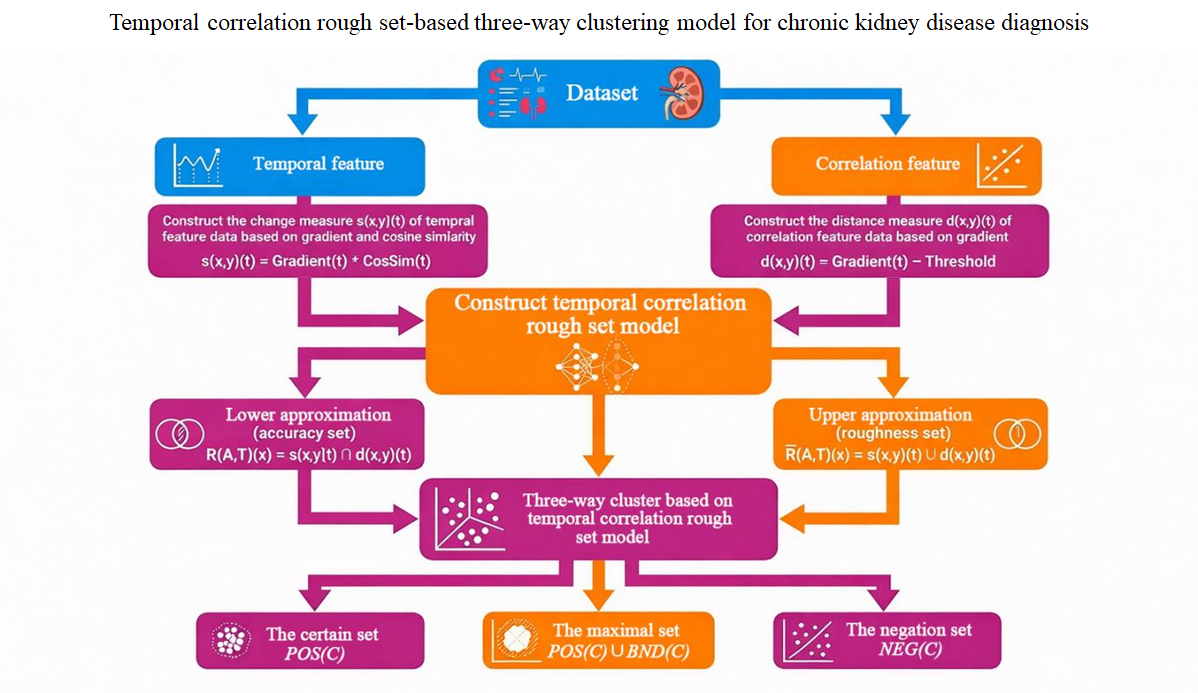
Chronic non-communicable diseases (CNCDs) are the leading cause of morbidity and mortality worldwide. Currently, diagnostic information for CNCDs remains incomplete and is subject to ongoing change, leading to uncertainty in diagnostic conclusions and severely impacting disease control. Therefore, it is necessary to study decision-making theory and methods for assisting clinical diagnosis. Given the characteristics of clinical diagnostic decision problems, this paper discusses the temporally correlated rough set theory and its three-way clustering model. Firstly, the temporal correlation rough set model based on the gradient and cosine similarity is defined. Then, the clinical diagnostic decision process for CNCDs is transformed into a three-way clustering problem with temporal correlation attributes. A three-way clustering model based on a temporal correlation rough set is constructed to support clinical diagnosis decisions for CNCDs. In this model, the common features of temporal association attributes are identified using a positive domain-based three-way clustering algorithm. Finally, the validity and applicability of the theoretical model are verified using real clinical data from 80,139 visit time points and 83 laboratory examination indicators from 3,094 chronic kidney disease (CKD) patients. Based on the algorithm’s calculations and a retrospective literature review, β2-microglobulin is identified as a candidate marker for CKD that warrants further validation and could provide auxiliary decision support for clinical practice. The main contribution of this paper is twofold. One is to make a new theoretical contribution to the data-driven research paradigm of clinical diagnosis decision-making. Another is to provide quantitative decision support for clinical diagnosis.
- Quam L, Smith R, Yach D. Rising to the global challenge of the chronic disease epidemic. Lancet. 2006;368(9543):1221–1223. doi: 10.1016/S0140-6736(06)69422-1
- Furman D, Campisi J, Verdin E, et al. Chronic inflammation in the etiology of disease across the life span. Nat Med. 2019;25(12):1822–1832. doi: 10.1038/s41591-019-0675-0
- Kalantar-Zadeh K, Li P. Strategies to prevent kidney disease and its progression. Nat Rev Nephrol. 2020;16(3):129–130. doi: 10.1038/s41581-020-0253-1
- GBD Chronic Kidney Disease Collaboration. Global, regional, and national burden of chronic kidney disease, 1990–2017: a systematic analysis for the Global Burden of Disease Study 2017. Lancet. 2020;395(10225):709–733. doi: 10.1016/s0140-6736(20)30045-3
- Feinstein A. The pre-therapeutic classification of co-morbidity in chronic disease. J Chronic Dis. 1970;23(7):455–468. doi: 10.1016/0021-9681(70)90054-8
- Stockley R, Halpin D, Celli B, et al. Chronic obstructive pulmonary disease biomarkers and their interpretation. Am J Respir Crit Care Med. 2019;199(9):1195–1204. doi: 10.1164/rccm.201810-1860SO
- Pinto-Plata V. Use of Proteomic Patterns of Serum Biomarkers in Patients with Chronic Obstructive Pulmonary Disease: Correlation with Clinical Parameters. Proc Am Thorac Soc. 2006;3(6):465-466. doi: 10.1513/pats.200603-030ms
- Sinha K, Uddin Z, Kawsar H, et al. Analyzing chronic disease biomarkers using electrochemical sensors and artificial neural networks. Trends Analyt Chem. 2023;158:116861. doi: 10.1016/j.trac.2022.116861
- Topalovic M, Laval S, Aerts JM, Troosters T, Decramer M, Janssens W. Automated Interpretation of Pulmonary Function Tests in Adults with Respiratory Complaints. Respiration. 2017;93(3):170-178. doi: 10.1159/000454956
- Liu Z, Liu L, Weng S, et al. Machine learning-based integration develops an immune-derived lncRNA signature for improving outcomes in colorectal cancer. Nat Commun. 2022;13(1):816. doi: 10.1038/s41467-022-28421-6
- Li J, Huang Q, Ren S, et al. A novel medical text classification model with Kalman filter for clinical decision making. Biomed Signal Process Control. 2023;82:104503. doi: 10.1016/j.bspc.2022.104503
- Choi MY, Chen I, Clarke AE, et al. Machine learning identifies clusters of longitudinal autoantibody profiles predictive of systemic lupus erythematosus disease outcomes. Ann Rheum Dis. 2023. doi: 10.1136/ard-2022-223808
- Wang X, Chen Y, Gao Y, et al. Predicting gastric cancer outcome from resected lymph node histopathology images using deep learning. Nat Commun. 2021;12(1):1637. doi: 10.1038/s41467-021-21674-7
- Pawlak Z. Rough sets. Int J Comput Inf Sci. 1982;11(5):341–356. doi: 10.1007/BF01001956
- Xing J, Gao C, Zhou J. Weighted fuzzy rough sets-based tri-training and its application to medical diagnosis. Appl Soft Comput. 2022;124:109025. doi: 10.1016/j.asoc.2022.109025
- Srivastava S, Meena YK, Singh G. Forecasting on COVID-19 infection waves using a rough set filter-driven moving average model. Appl Soft Comput. 2022;131:109750. doi: 10.1016/j.asoc.2022.109750
- Tsumoto S. Mining diagnostic rules from clinical databases using rough sets and medical diagnostic model. Inf Sci. 2004;162(2):65–80. doi: 10.1016/j.ins.2004.03.002
- Yao Y. Two views of the theory of rough sets in finite universes. Int J Approx Reason. 1996;15(4):291–317. doi: 10.1016/S0888-613X(96)00071-0
- Grzymala-Busse JW. LERS-A System for Learning from Examples Based on Rough Sets. In: Intelligent Decision Support. Springer; 1992:3-18. doi: 10.1007/978-94-015-7975-9_1
- Li W, Yang B. Three-way decisions with fuzzy probabilistic covering-based rough sets and their applications in credit evaluation. Appl Soft Comput. 2023;136:110144. doi: 10.1016/j.asoc.2023.110144
- Qahtan S, Alsattar HA, Zaidan AA, Deveci M, Pamucar D, Delen D. Performance assessment of sustainable transportation in the shipping industry using a q-rung ortho-pair fuzzy rough sets-based decision making methodology. Expert Syst Appl. 2023;223:119958. doi: 10.1016/j.eswa.2023.119958
- Zheng Q, Chang H, Liu Z, et al. Multi-stage design space reduction technology based on SOM and rough sets, and its application to hull form optimization. Expert Syst Appl. 2023;213:119229. doi: 10.1016/j.eswa.2022.119229
- Bai J, Guo J, Sun B, et al. Intelligent forecasting model of stock price using neighborhood rough set and multivariate empirical mode decomposition. Eng Appl Artif Intell. 2023;122:106106. doi: 10.1016/j.engappai.2023.106106
- Tishya M, Anitha A. Precipitation prediction by integrating rough set on fuzzy approximation space with deep learning techniques. Appl Soft Comput. 2023;139:110253. doi: 10.1016/j.asoc.2023.110253
- Yan HY, Zhang XR, Dong JH, Shang MS, Shan K, Wu D, et al. Spatial and temporal relation rule acquisition of eutrophication in Da’ning River based on rough set theory. Ecol Indic. 2016;66:180-189. doi: 10.1016/j.ecolind.2016.01.032
- Yao Y .The superiority of three-way decisions in probabilistic rough set models. Inf Sci. 2011;181(6):1080-1096. doi: 10.1016/j.ins.2010.11.019.
- Podsiadlo M , Rybinski H .Financial Time Series Forecasting using Rough Sets with Time-Weighted Rule Voting. Expert Systems with Applications, 2016;66:219-233. doi: 10.1016/j.eswa.2016.08.066.
- Selvakumar K, Karuppiah M, SaiRamesh L, Islam SH, Hassan MM, Fortino G, et al. Intelligent temporal classification and fuzzy rough set-based feature selection algorithm for intrusion detection system in WSNs. Inf Sci. 2019;497:77-90. doi: 10.1016/j.ins.2019.05.040
- Dong L, Wang R, Chen D. Incremental feature selection with fuzzy rough sets for dynamic data sets. Fuzzy Sets Syst. 2023;467:108503. doi: 10.1016/j.fss.2023.03.006
- Ni P, Zhao S, Wang X. Incremental feature selection based on fuzzy rough sets. Inf Sci. 2020;536:185–204. doi: 10.1016/j.ins.2020.04.038
- Che X, Chen D, Mi J. Label correlation in multi-label classification using local attribute reductions with fuzzy rough sets. Fuzzy Sets Syst. 2022;426:121–144. doi: 10.1016/j.fss.2021.03.016
- Lee S, Enke D, Kim Y. A relative value trading system based on a correlation and rough set analysis for the foreign exchange futures market. Eng Appl Artif Intell. 2017;61:47-56. doi: 10.1016/j.engappai.2017.02.014
- Afridi MK, Azam N, Yao J. Variance based three-way clustering approaches for handling overlapping clustering. Int J Approx Reason. 2020;118:47-63. doi: 10.1016/j.ijar.2019.11.011
- Yu H, Chen L, Yao J. A three-way clustering method based on an improved DBSCAN algorithm. Phys A. 2019;535:122289. doi: 10.1016/j.physa.2019.122289
- Sun C, Du M, Sun J, Li K, Dong Y. A three-way clustering method based on improved density peaks algorithm and boundary detection graph. Int J Approx Reason. 2023;153:239-257. doi: 10.1016/j.ijar.2022.12.002
- Yu H, Chen L, Yao J. A three-way density peak clustering method based on evidence theory. Knowl Based Syst. 2021;211:106532. doi: 10.1016/j.knosys.2020.106532
- Ali B, Azam N, Yao J. A three-way clustering approach using image enhancement operations. Int J Approx Reason. 2022;149:1–38. doi: 10.1016/j.ijar.2022.07.001
- Yu H, Zhang C, Wang G. A tree-based incremental overlapping clustering method using three-way decision theory. Knowl Based Syst. 2016;91:189–203. doi: 10.1016/j.knosys.2015.05.028
- Yao Y. Three-way decision and granular computing. Int J Approx Reason. 2018;103:107–123. doi: 10.1016/j.ijar.2018.09.005
- Cha S, Shin I, Kim D, et al. Effectiveness of serum β2-microglobulin for evaluating donor kidney status. Sci Rep. 2020;10:8109. doi: 10.1038/s41598-020-65134-6
- Barton KT, Kakajiwala A, Dietzen DJ, Goss CW, Gu H, Dharnidharka VR. Using the newer Kidney Disease: Improving Global Outcomes criteria, beta-2-microglobulin levels associate with severity of acute kidney injury. Clin Kidney J. 2018;11(6):797-802. doi: 10.1093/ckj/sfy056
- Kanda E, Muenz D, Bieber B, Cases A, Locatelli F, Port FK, et al. Beta-2 microglobulin and all-cause mortality in the era of high-flux hemodialysis: results from the Dialysis Outcomes and Practice Patterns Study. Clin Kidney J. 2020;14(5):1436-1442. doi: 10.1093/ckj/sfaa155
- Gejyo F, Yamada T, Odani S, Nakagawa Y, Arakawa M, Kunitomo T, et al. A new form of amyloid protein associated with chronic hemodialysis was identified as β2-microglobulin. Biochem Biophys Res Commun. 1985;129(3):701-706. doi: 10.1016/0006-291x(85)91948-5
- Fishbane S, Kalantar‐Zadeh K, Nissenson AR. Serum Ferritin in Chronic Kidney Disease: Reconsidering the Upper Limit for Iron Treatment. Semin Dial. 2004;17(5):336-341. doi: 10.1111/j.0894-0959.2004.17359.x
- McCullough K, Bolisetty S. Iron homeostasis and ferritin in sepsis-associated kidney injury. Nephron. 2020;144(12):616–620. doi: 10.1159/000509073
- Fleming S. Immunocytochemical Localization of Ferritin in the Kidney and Renal Tumours. Eur Urol. 1987;13(6):407-411. doi: 10.1159/000472835
- Abrahamson DR, John PLSt. Ultrastructure of developing kidney glomerular basement membranes: Temporal changes in binding of anti‐laminin lgG and cationized ferritin. Microsc Res Tech. 1994;28(2):81-94. doi: 10.1002/jemt.1070280202
- Operskalski J, Barbey A. Risk literacy in medical decision-making. Science. 2016;352(6284):413–414. doi: 10.1126/science.aaf7966
- Athey S. Beyond prediction: Using big data for policy problems. Science. 2017;355(6324):483–485. doi: 10.1126/science.aal4321
- Meara S. China’s data-driven dream to overhaul health care. Nature. 2021;598:S1–S3. doi: 10.1038/d41586-021-02694-1

